23andme-My Haplogroup mtDNA L2a1c1
This is a printable version of your report and may contain sensitive and personalized information.
Please when you are done printing.

ANTONIO FLORENTINO
Maternal Haplogroup
You descend from a long line of women that can be traced back to eastern Africa over 150,000 years ago. These are the women of your maternal line, and your maternal haplogroup sheds light on their story.
ANTONIO, your maternal haplogroup is L2a1c1.
As our ancestors ventured out of eastern Africa, they branched off in diverse groups that crossed and recrossed the globe over tens of thousands of years. Some of their migrations can be traced through haplogroups, families of lineages that descend from a common ancestor. Your maternal haplogroup can reveal the path followed by the women of your maternal line.
Migrations of Your Maternal Line

Haplogroup L
If every person living today could trace his or her maternal line back over thousands of generations, all of our lines would meet at a single woman who lived in eastern Africa between 150,000 and 200,000 years ago. Though she was one of perhaps thousands of women alive at the time, only the diverse branches of her haplogroup have survived to today. The story of your maternal line begins with her.
Haplogroup L2
Your branch of L is haplogroup L2, which arose from a woman who lived in Africa almost 90,000 years ago. More recently — in fact, only 4,000 years ago — her descendants were a part of the major Bantu migrations which carried them from central Africa to the east and south and made L2 the most common haplogroup among Africans.
Origin and Migrations of Haplogroup L2a1
Your maternal line stems from a branch of haplogroup L2 called L2a1. L2a1 is a widespread branch that arose approximately 24,000 years ago. At the peak of the Last Ice Age 20,000 years ago, the Sahara Desert became entirely uninhabitable and began expanding to the south. People in central Africa bearing L2a1 began journeying in two directions: east toward the cooler climate of the eastern highlands, and west towards the Atlantic coast.
Then, beginning about 4,000 years ago, two sub-branches of L2a1 — L2a1a and L2a1b — were swept up with the Bantu-speaking people of West Africa. These people, who had been practicing farming for a thousand years or more, began to expand their territory and gradually introduced both their language and their way of life to their eastern and southern neighbors. Today, both L2a1a and L2a1b are well-represented in Southeast Africa.
L2a1 is also quite common among the descendants of Africans in North and South America. Its high frequency is probably due to the concentration of L2a1 in West Africa, which was the main supply region for the Atlantic slave trade.
L2a1c1
Your maternal haplogroup, L2a1c1, traces back to a woman who lived approximately 7,000 years ago.
L2a1c1 is relatively uncommon among 23andMe customers.
The Bantu Expansion transformed the cultural and genetic landscape of Africa.
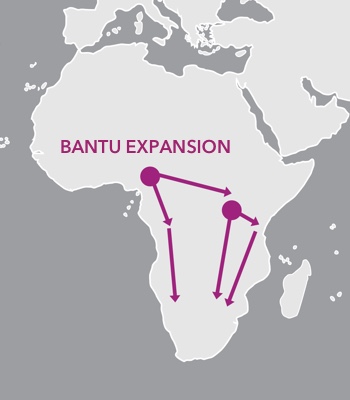
About 5,000 years ago, many people in sub-Saharan Africa still relied on hunting, gathering, and foraging as their main means of collecting food. But that was soon going to change. People in West-Central Africa began experimenting with agriculture, cultivating the yams, legumes, peppers, and gourds that would became staples of sub-Saharan African diet. These people spoke languages belonging to the Bantu language family, and about 4,000 years ago they began to move.
First, they headed east across the central rainforest. Eventually, the descendants of these migrants arrived at the farthest reaches of southern Africa. Later, other Bantu speakers who had remained in West Africa also began to travel down the western coast. As they traveled over a period of centuries, they both displaced and absorbed many other hunter-gatherer groups that were already living throughout Africa.Their agricultural and technological knowledge also diffused to other local groups. They often intermarried, sometimes adopting local cultural practices of those people they encountered. The languages that they brought with them from their ancestral homeland spread throughout sub-Saharan Africa, and today the majority of sub-Saharan African languages are Bantu.
The Genetics of Maternal Haplogroups
Scientific Details
Your haplogroup is determined by your mitochondrial DNA.
Each generation, mothers pass down copies of their mitochondrial DNA (mtDNA) to their children. While most of your genome exists in 23 pairs of chromosomes that exchange pieces between generations in a process called recombination, mtDNA is transmitted unshuffled. Because of this unusual pattern of inheritance, mtDNA contains rich information about maternal lineages.
A small number of DNA changes, called mutations, generally occur from one generation to the next. Because mtDNA does not recombine between generations, these mutations accumulate in patterns that uniquely mark individual lineages. Scientists can compare the sequence differences that result by constructing a tree. This tree shows how maternal lineages relate to one another, including the observation that they all share a most recent common ancestor approximately 150,000 to 200,000 years ago.
The term "haplogroup" refers to a family of lineages that share a common ancestor and, therefore, a particular set of mutations. We identify your haplogroup by determining which branches of the mtDNA tree correspond to your DNA. Because more closely related lineages tend to share geographic roots, your haplogroup can provide insight into the origins of some of your ancient maternal-line ancestors.
Maternal haplogroups are named with sequences of letters and numbers that reflect the structure of the tree and how the branches relate to one another.
References
- Abu-Amero KK et al. (2007). "Eurasian and African mitochondrial DNA influences in the Saudi Arabian population." BMC Evol Biol. 7:32.
- Barbieri C et al. (2014). "Migration and interaction in a contact zone: mtDNA variation among Bantu-speakers in Southern Africa." PLoS One. 9(6):e99117.
- Behar DM et al. (2012). "A "Copernican" reassessment of the human mitochondrial DNA tree from its root." Am J Hum Genet. 90(4):675-84.
- Beleza S et al. (2005). "The genetic legacy of western Bantu migrations." Hum Genet. 117(4):366-75.
- Bodner M et al. (2012). "Rapid coastal spread of First Americans: novel insights from South America's Southern Cone mitochondrial genomes." Genome Res. 22(5):811-20.
- Brandstätter A et al. (2004). "Mitochondrial DNA control region sequences from Nairobi (Kenya): inferring phylogenetic parameters for the establishment of a forensic database." Int J Legal Med. 118(5):294-306.
- Cerný V et al. (2004). "mtDNA sequences of Chadic-speaking populations from northern Cameroon suggest their affinities with eastern Africa." Ann Hum Biol. 31(5):554-69.
- Cerný V et al. (2006). "MtDNA of Fulani nomads and their genetic relationships to neighboring sedentary populations." Hum Biol. 78(1):9-27.
- Cerný V et al. (2007). "A bidirectional corridor in the Sahel-Sudan belt and the distinctive features of the Chad Basin populations: a history revealed by the mitochondrial DNA genome." Ann Hum Genet. 71(Pt 4):433-52.
- Chan EK et al. (2015). "Revised timeline and distribution of the earliest diverged human maternal lineages in southern Africa." PLoS One. 10(3):e0121223.
- Coia V et al. (2005). "Brief communication: mtDNA variation in North Cameroon: lack of Asian lineages and implications for back migration from Asia to sub-Saharan Africa." Am J Phys Anthropol. 128(3):678-81.
- Destro-Bisol G et al. (2004). "The analysis of variation of mtDNA hypervariable region 1 suggests that Eastern and Western Pygmies diverged before the Bantu expansion." Am Nat. 163(2):212-26.
- Ely B et al. (2006). "African-American mitochondrial DNAs often match mtDNAs found in multiple African ethnic groups." BMC Biol. 4:34.
- Fadhlaoui-Zid K et al. (2004). "Mitochondrial DNA heterogeneity in Tunisian Berbers." Ann Hum Genet. 68(Pt 3):222-33.
- Fernandes V et al. (2012). "The Arabian cradle: mitochondrial relicts of the first steps along the southern route out of Africa." Am J Hum Genet. 90(2):347-55.
- Fu Q et al. (2016). "The genetic history of Ice Age Europe." Nature. 534(7606):200-5.
- González AM et al. (2003). "Mitochondrial DNA affinities at the Atlantic fringe of Europe." Am J Phys Anthropol. 120(4):391-404.
- Jackson BA et al. (2005). "Mitochondrial DNA genetic diversity among four ethnic groups in Sierra Leone." Am J Phys Anthropol. 128(1):156-63.
- Kayser M. (2010). "The human genetic history of Oceania: near and remote views of dispersal." Curr Biol. 20(4):R194-201.
- Kivisild T et al. (2004). "Ethiopian mitochondrial DNA heritage: tracking gene flow across and around the gate of tears." Am J Hum Genet. 75(5):752-70.
- López S et al. (2015). "Human Dispersal Out of Africa: A Lasting Debate." Evol Bioinform Online. 11(Suppl 2):57-68.
- Llamas B et al. (2016). "Ancient mitochondrial DNA provides high-resolution time scale of the peopling of the Americas." Sci Adv. 2(4):e1501385.
- Majumder PP. (2010). "The human genetic history of South Asia." Curr Biol. 20(4):R184-7.
- Malaspinas AS et al. (2016). "A genomic history of Aboriginal Australia." Nature. 538(7624):207-214.
- Mellars P et al. (2013). "Genetic and archaeological perspectives on the initial modern human colonization of southern Asia." Proc Natl Acad Sci U S A. 110(26):10699-704.
- Metspalu M et al. (2004). "Most of the extant mtDNA boundaries in south and southwest Asia were likely shaped during the initial settlement of Eurasia by anatomically modern humans." BMC Genet. 5:26.
- Plaza S et al. (2003). "Joining the pillars of Hercules: mtDNA sequences show multidirectional gene flow in the western Mediterranean." Ann Hum Genet. 67(Pt 4):312-28.
- Quintana-Murci L et al. (2004). "Where west meets east: the complex mtDNA landscape of the southwest and Central Asian corridor." Am J Hum Genet. 74(5):827-45.
- Rando JC et al. (1998). "Mitochondrial DNA analysis of northwest African populations reveals genetic exchanges with European, near-eastern, and sub-Saharan populations." Ann Hum Genet. 62(Pt 6):531-50.
- Richards M et al. (2000). "Tracing European founder lineages in the Near Eastern mtDNA pool." Am J Hum Genet. 67(5):1251-76.
- Rito T et al. (2013). "The first modern human dispersals across Africa." PLoS One. 8(11):e80031.
- Rosa A et al. (2004). "MtDNA profile of West Africa Guineans: towards a better understanding of the Senegambia region." Ann Hum Genet. 68(Pt 4):340-52.
- Soares P et al. (2009). "Correcting for purifying selection: an improved human mitochondrial molecular clock." Am J Hum Genet. 84(6):740-59.
- Stoneking M et al. (2010). "The human genetic history of East Asia: weaving a complex tapestry." Curr Biol. 20(4):R188-93.
- Tishkoff SA et al. (2007). "History of click-speaking populations of Africa inferred from mtDNA and Y chromosome genetic variation." Mol Biol Evol. 24(10):2180-95.
Change Log
Your report may occasionally be updated based on new information. This Change Log describes updates and revisions to this report.
| Date | Change |
|---|---|
| May 8, 2017 | The standalone Maternal Haplogroup report was created, featuring new design elements and content. |
| Oct. 21, 2015 | Haplogroups report created. |







Comentários
Postar um comentário